In temperate forests, leaf phenology is a sensitive indicator of climate change and a major regulator of seasonal carbon and water cycling. Intra-site leaf phenology variability across individual trees and species has been widely observed, but traditional approaches for monitoring individual tree-scale leaf phenology are often limited to a small spatial extent and sample size. Recent availability of PlanetScope satellite data with a 3 m spatial resolution, near-daily revisiting frequency, and global coverage provides opportunities to overcome this limitation. However, comprehensive assessments of PlanetScope’s capacity and scalability for fine-scale leaf phenology monitoring remain lacking.
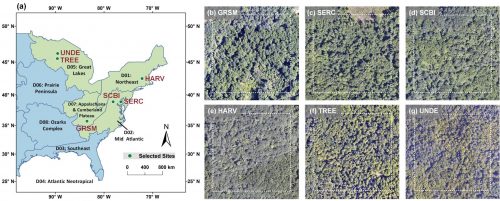
An international research team of scientists from Hong Kong, Mainland China, the United States, and Brazil, led by Dr. Jin Wu (Assistant Professor from the School of Biological Sciences, HKU), including Ms. Yingyi Zhao (PhD candidate from the School of Biological Sciences, HKU), Dr. Calvin K.F. Lee (Postdoctoral fellow from the School of Biological Sciences, HKU), and Dr. Timothy Bonebrake (Associate Professor from the School of Biological Sciences, HKU ), have proposed an approach to comprehensively evaluate PlanetScope’s capacity for individual tree/species-scale leaf phenology monitoring. This approach integrates 0.1 m resolution airborne imagery and ground phenology records of individual trees with 3m PlanetScope image time series. With this approach, the team used data from six NEON forest sites in eastern North America and extracted key phenological metrics at the individual tree scale and species scale respectively from PlanetScope satellites and ground observations over 2018 and 2019. The capacity of PlanetScope satellites for fine-scale phenology monitoring was evaluated by comparing PlanetScope-derived individual tree/species-scale phenology with corresponding ground data.
According to the results of assessments, the team found that PlanetScope-derived fine-scale land surface characterize significant leaf phenology variability at the individual tree scale across all forest sites and years. The accuracy was improved at the species level when more PlanetScope pixels were included; and 2) captured relatively more variation in fall phenology relative to spring phenology. In addition, PlanetScope satellites were also superior to other conventional means (e.g., satellites with coarse resolution, ground observations, and PhenoCam observations) in providing landscape-scale phenology information with spatially explicit details.
“Our study presents a comprehensive evaluation of PlanetScope’s capacity for individual tree-scale and species-scale leaf phenology monitoring and highlights the potential of PlanetScope to provide rich fine-scale phenology information to help advance the field of plant phenology research,” said Dr. Jin Wu, Principal Investigator of Global Ecology and Remote Sensing (GEARS) lab at HKU. Ms. Yingyi Zhao added “In this study, we found that there is significant phenology variation existing among different individual trees and species based on the PlanetScope-derived fine-scale land surface phenology (LSP). Therefore, PlanetScope-derived fine-scale LSP characterizations over large vegetated landscapes might generate new opportunities across various levels from individual trees to forest ecosystems. Moreover, if longer-term observations from PlanetScope and other CubeSat constellations become available, we will have rich fine-scale phenology information over consecutive years and this data potentially could advance our understanding of the phenological responses to the changing climate across various scales”.
In the next step, the GEARS lab is aiming to leverage fine-scale phenology information provided by CubeSats and other satellite data to further advance our understanding of phenology-related questions, such as the underlying controls of fine-scale phenology variability and the impacts of fine-scale phenology variability on the plant carbon uptake.
The findings were recently published in Remote Sensing of Environment.
The journal paper can be accessed from here